jupyter-cytoscape
v1.3.3
Published
A Cytoscape Jupyter Widget
Downloads
298
Readme
ipycytoscape
A widget enabling interactive graph visualization with cytoscape.js in JupyterLab and the Jupyter notebook.
Try it out using binder: or install and try out the examples.
Supports:
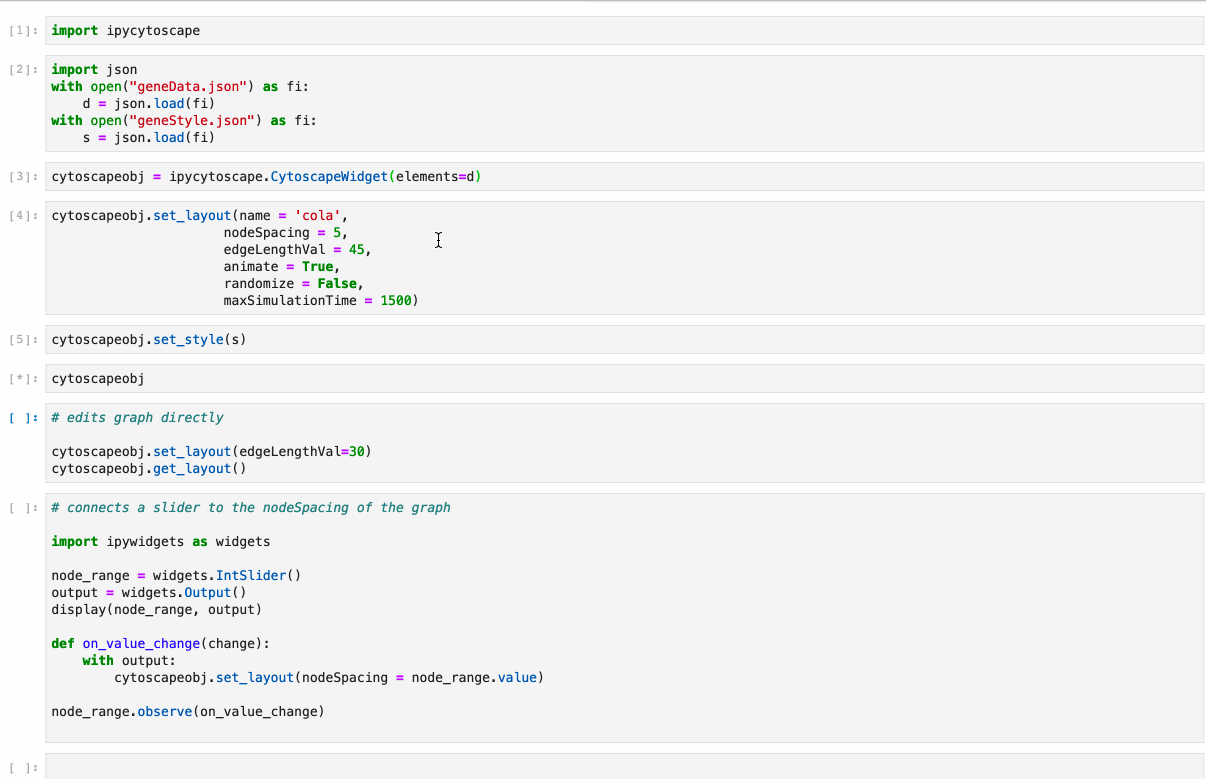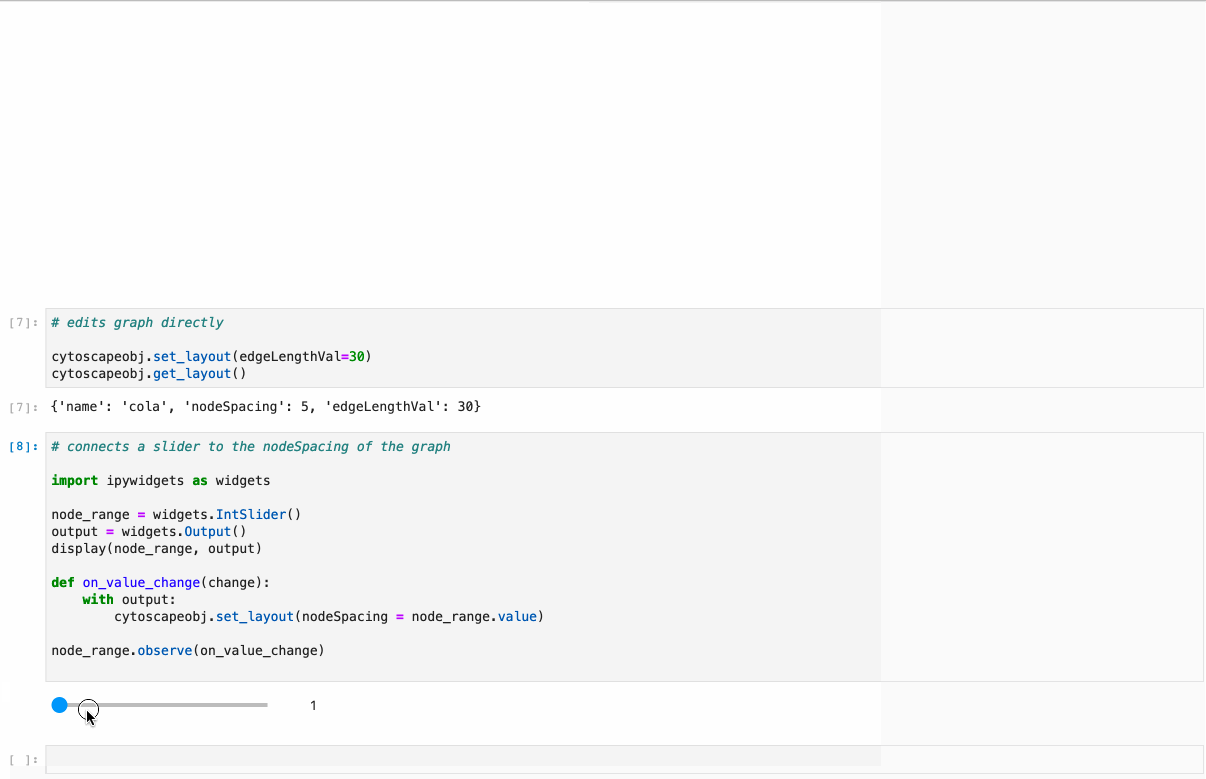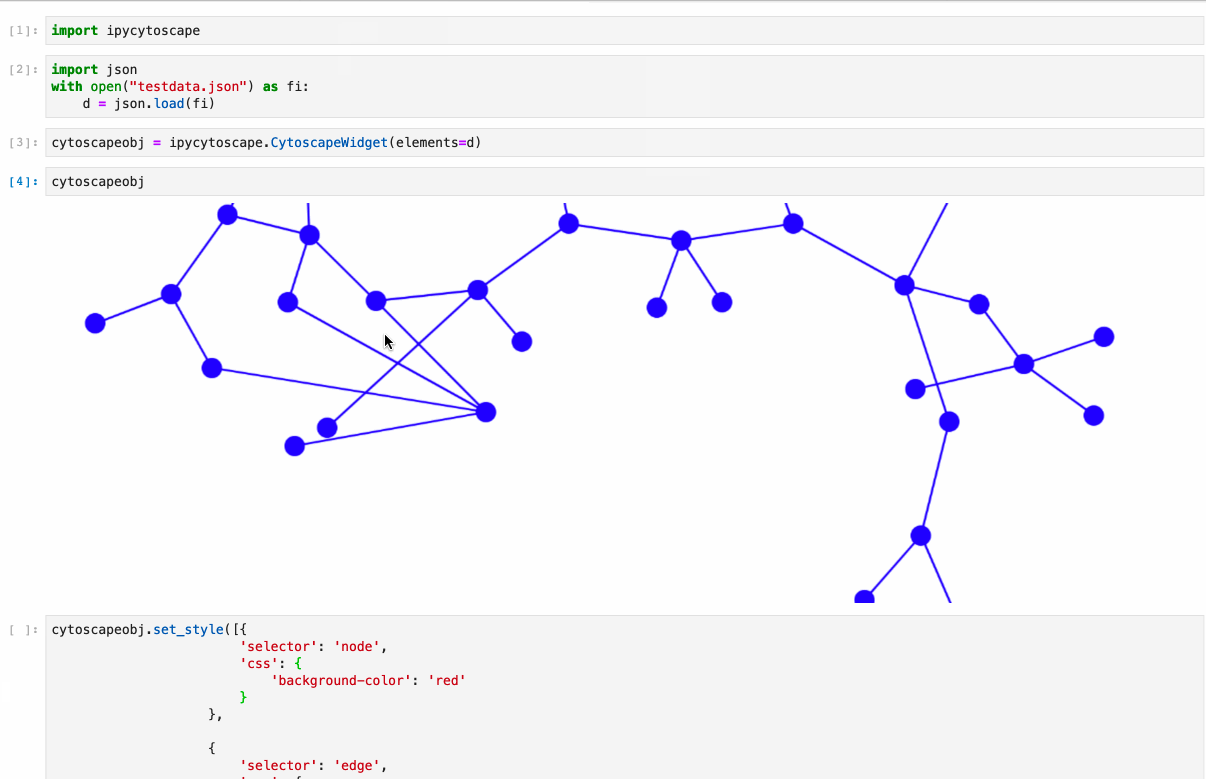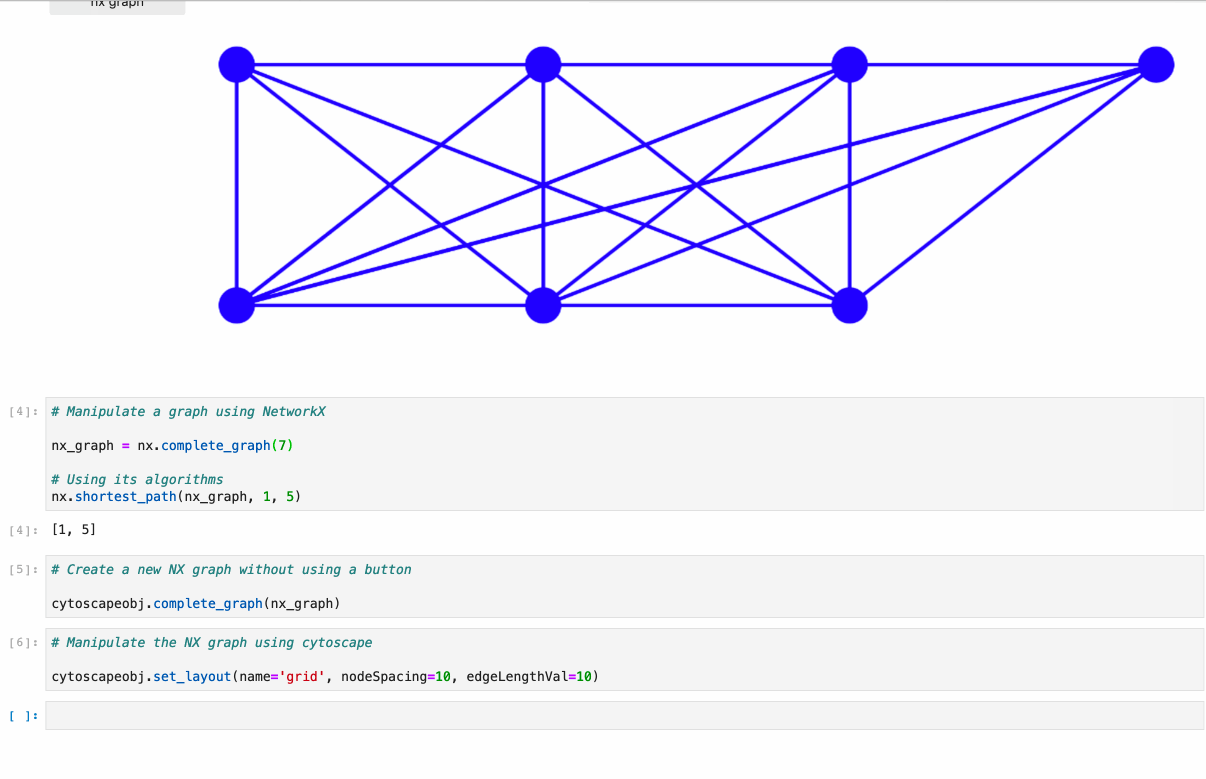
- Conversion from NetworkX see example1, example2
- Conversion from Pandas DataFrame see example
- Conversion from neo4j see example
Installation
With mamba:
mamba install -c conda-forge ipycytoscapeWith conda:
conda install -c conda-forge ipycytoscapeWith pip:
pip install ipycytoscapePandas installation
You can install the Pandas dependencies for ipycytoscape with pip:
pip install pandasOr conda-forge:
mamba install pandasNeo4j installation
You can install the neo4j dependencies for ipycytoscape with pip:
pip install -e ".[neo4j]"Or conda-forge:
mamba install py2neo neotimeFor jupyterlab 1.x or 2.x:
If you are using JupyterLab 1.x or 2.x then you will also need to install nodejs and the jupyterlab-manager extension. You can do this like so:
# installing nodejs
conda install -c conda-forge nodejs
# install jupyterlab-manager extension
jupyter labextension install @jupyter-widgets/[email protected] --no-build
# if you have previously installed the manager you still to run jupyter lab build
jupyter lab buildFor Jupyter Notebook 5.2 and earlier
You may also need to manually enable the nbextension:
jupyter nbextension enable --py [--sys-prefix|--user|--system] ipycytoscapeFor a development installation:
(requires npm)
While not required, we recommend creating a conda environment to work in:
conda create -n ipycytoscape -c conda-forge jupyterlab nodejs
conda activate ipycytoscape
# clone repo
git clone https://github.com/cytoscape/ipycytoscape.git
cd ipycytoscapeInstall python package for development
This will run npm install and npm run build.
This command will also install the test suite and the docs locally:
pip install jupyter_packaging==0.7.9
pip install -e ".[test, docs]"
jupyter labextension develop . --overwriteoptionally install the pre-commit hooks with:
pre-commit installOr for classic notebook, you can run:
jupyter nbextension install --sys-prefix --symlink --overwrite --py ipycytoscape
jupyter nbextension enable --sys-prefix --py ipycytoscapeNote that the --symlink flag doesn't work on Windows, so you will here have to run
the install command every time that you rebuild your extension. For certain installations
you might also need another flag instead of --sys-prefix, but we won't cover the meaning
of those flags here.
How to see your changes
Typescript:
To continuously monitor the project for changes and automatically trigger a rebuild, start watching the ipycytoscape code:
npm run watchAnd in a separate terminal start JupyterLab normally:
jupyter labonce the webpack rebuild finishes refresh the JupyterLab page to have your changes take effect.
Python:
If you make a change to the python code then you need to restart the notebook kernel to have it take effect.
How to run tests locally
Install necessary dependencies with pip:
pip install -e ".[test]"Or with conda/mamba:
mamba -c conda-forge install networkx pandas nbval pytestAnd to run it:
pytestHow to build the docs
cd docs
Install dependencies:
conda env update --file doc_environment.yml
And build them:
make html
Acknowledgements
The ipycytoscape project was started by Mariana Meireles at QuantStack. This initial development was funded as part of the PLASMA project, which is led by Claire Vandiedonck, Pierre Poulain, and Sandrine Caburet.
License
We use a shared copyright model that enables all contributors to maintain the copyright on their contributions.
This software is licensed under the BSD-3-Clause license. See the LICENSE file for details.

